What is Multilabel Classification?
Multilabel classification is a type of supervised learning where each sample can be assigned multiple labels simultaneously, rather than being restricted to a single category. Unlike traditional multiclass classification, which forces a model to choose one class from many options, multilabel classification allows images to belong to multiple classes at once.
For example, in a clothing classification system:
- Multiclass Classification would categorize an image as either "t-shirt" OR "jeans" OR "shoes"
- Multilabel Classification could recognize "t-shirt" AND "blue" AND "cotton" AND "casual" all in the same image
This approach is particularly valuable when working with complex visual data where multiple attributes, objects, or characteristics coexist within a single frame. Each label is treated as an independent binary decision, allowing the model to learn which combinations of labels are likely to appear together.
Key Differences with Single Class Annotation
Understanding how multilabel classification differs from traditional single-class annotation is crucial for effective implementation:
Label Independence: In multiclass classification, labels are mutually exclusive - an image can only belong to one category. In multilabel classification, each label is independent, meaning an image can have zero, one, or multiple labels applied simultaneously.
Annotation Flexibility: Single-class annotation requires you to choose the "best fit" category for each image, even when multiple characteristics are present. Multilabel annotation allows you to capture all relevant attributes without compromise, providing a more comprehensive representation of your data.
Model Output Interpretation: Multiclass models output probabilities that sum to 1 across all classes, essentially answering "which one category best describes this image?" Multilabel models output independent probabilities for each label, answering "which of these labels apply to this image?" Each probability is evaluated separately.
Training Data Richness: Multilabel datasets capture more nuanced information about your images. Instead of forcing images into single categories, you can represent the full complexity of your visual data, leading to models that understand multiple dimensions of your problem space.
Use Case Alignment: Choose multiclass when images naturally fall into distinct, non-overlapping categories (e.g., animal species identification). Choose multilabel when images can have multiple simultaneous properties (e.g., image tagging, attribute recognition, or multi-property analysis).
.gif)
Training Losses Compared
The choice of loss function is fundamental to how a classification model learns. The difference between multiclass and multilabel classification is reflected in their respective loss functions. We explain the following based on these variables:

.svg)
.svg)
.svg)
Classification (Multiclass) - Cross Entropy Loss
Standard multiclass classification on Datature uses cross entropy loss, which is designed for mutually exclusive categories. This loss function:
- Applies a softmax activation to produce probabilities that sum to 1 across all classes
- Penalizes the model based on the predicted probability of the true class
- Works on the assumption that exactly one label is correct for each image
The formula is as follows:
.svg)
Multilabel Classification - Binary Cross Entropy with Logits
Multilabel classification uses binary cross entropy with logits, which treats each label as an independent binary classification problem. This loss function:
- Applies a sigmoid activation to each label independently, producing separate probabilities that don't need to sum to 1
- Calculates loss for each label separately and averages them
- Allows multiple labels to be true (or false) simultaneously for the same image
- Combines the sigmoid activation and binary cross entropy into a single, numerically stable operation
The formula is as follows:
.svg)
The "with logits" aspect means the loss function operates directly on the raw model outputs before sigmoid activation, improving numerical stability and computational efficiency. This is a subtle but important implementation detail that prevents potential numerical precision issues when probabilities are very close to 0 or 1.
In practice, this means your multilabel model learns to make independent decisions for each label, rather than competing decisions across labels. This architectural difference is what enables the model to recognize multiple attributes simultaneously.
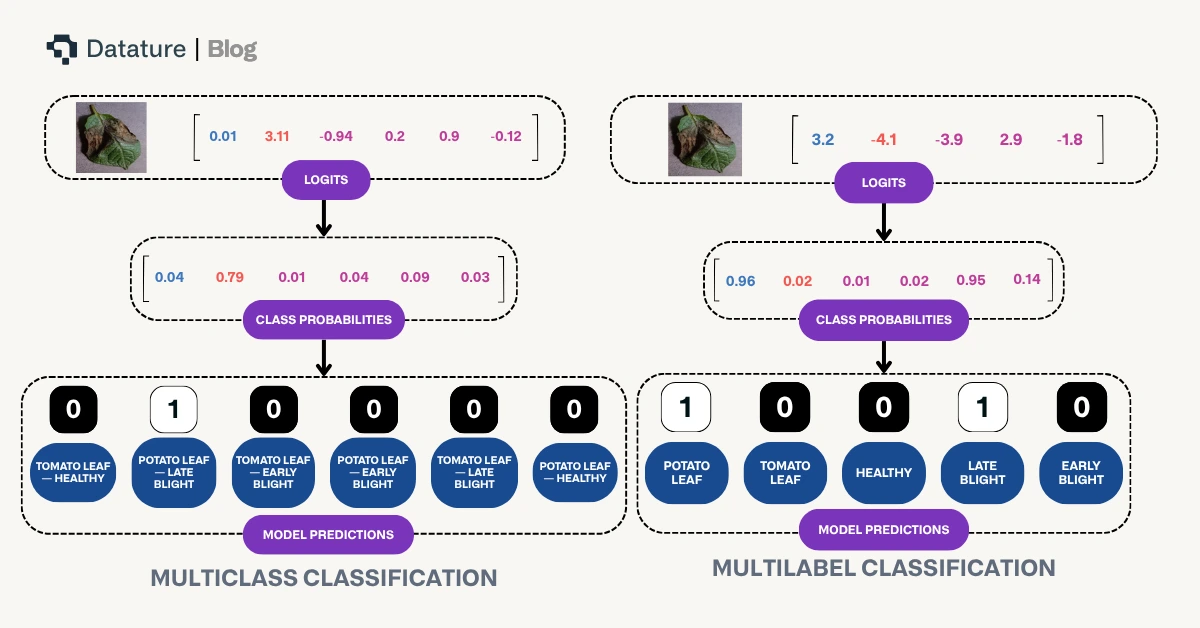
The above diagram highlights differences in how model prediction occurs based on these two differing paradigms.
Evaluating Model Performance: From Confusion Matrix to Co-occurrence Matrix
Understanding how to evaluate your model's performance is crucial, and the evaluation methods differ between multiclass and multilabel classification:
Confusion Matrix in Multiclass Classification
For traditional multiclass classification on Datature, the confusion matrix has been the primary tool for analyzing model performance. It displays predicted classes against actual classes in a grid format, making it easy to identify which categories the model confuses with one another. Each image contributes to exactly one cell in the matrix since it has a single true label and a single predicted label.
.webp)
Co-occurrence Matrix in Multilabel Classification
For multilabel classification, the confusion matrix concept doesn't translate directly because each image can have multiple true labels and multiple predicted labels. Instead, Datature uses a co-occurrence matrix to visualize label relationships and patterns. While these matrices look similar, they serve fundamentally different purposes.
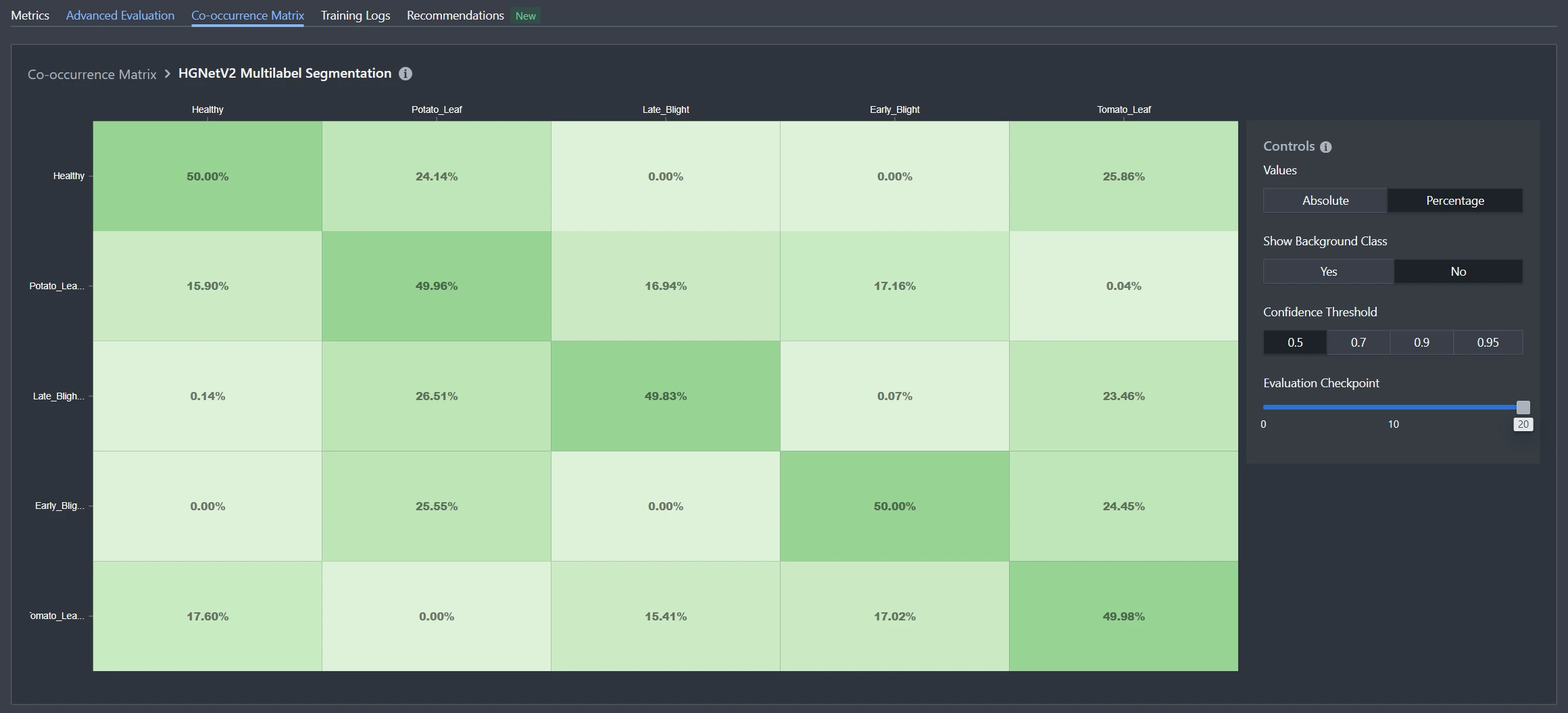
Similarities Between the Two Matrices
Both matrices provide:
- A grid-based visual representation of label relationships
- Numerical values that indicate the strength of relationships between labels
- An intuitive way to identify patterns at a glance
- Actionable insights for improving model performance or dataset quality
Key Differences
The fundamental differences reflect the nature of each classification type:
Purpose: A confusion matrix shows prediction errors and model accuracy by comparing predicted vs actual classes. A co-occurrence matrix shows which labels tend to appear together in your dataset or predictions, regardless of correctness.
Structure: In a confusion matrix, rows represent actual classes and columns represent predicted classes (or vice versa), with diagonal values indicating correct predictions. In a co-occurrence matrix, both axes represent the same set of labels, showing how frequently label pairs appear together.
Values: Confusion matrix cells count how many times class A was predicted when class B was the true label. Co-occurrence matrix cells count how many images contain both label A and label B simultaneously.
Diagonal Meaning: In a confusion matrix, diagonal values (matching actual and predicted) represent correct classifications - the higher, the better. In a co-occurrence matrix, diagonal values represent how often a label appears by itself, which is simply informative rather than evaluative.
What We Can Observe from the Co-occurrence Matrix
The co-occurrence matrix reveals valuable insights about your multilabel dataset and model behavior:
Label Relationships: Quickly identify which labels frequently appear together. For example, in a clothing dataset, you might observe that "cotton" and "casual" often co-occur, or in a content moderation system, certain policy violations might cluster together.
Dataset Balance: Spot imbalances in label combinations. If certain label pairs never or rarely appear together in your training data, your model may struggle to predict those combinations even if they're valid in real-world scenarios.
Model Behavior Patterns: Compare co-occurrence matrices between your training set and your model's predictions. Significant differences can indicate whether your model has learned the underlying relationships correctly or is over- or under-predicting certain label combinations.
Data Quality Issues: Anomalous co-occurrence patterns can reveal annotation inconsistencies. For instance, if two labels should logically appear together but rarely do in your matrix, it might indicate labeling errors or missing annotations.
Feature Correlation: Strong co-occurrence between certain labels can inform decisions about feature engineering, data collection priorities, or even whether certain labels should be merged or split for your specific use case.
By analyzing the co-occurrence matrix alongside traditional metrics like per-label precision and recall, you gain a comprehensive understanding of how your multilabel classification model performs and how your dataset is structured.
Creating a Multilabel Classification Project On Nexus
Use Case: Agricultural Leaf Health Monitoring
Let's walk through building a multilabel classification project using a real-world example from the agricultural industry. Imagine you're an agricultural specialist who needs to identify both the type of crop and its health condition from leaf images. In this scenario, each leaf image needs to be classified across two dimensions simultaneously: the crop type (Potato or Tomato) and the health status (Healthy, Early Blight, or Late Blight). This is a perfect use case for multilabel classification because a single image must receive labels from both categories - for example, a leaf might be classified as both "Tomato" AND "Early Blight" at the same time.
Label Structure:
- Crop Type: Potato, Tomato
- Health Status: Healthy, Early Blight, Late Blight
Unlike traditional single-class classification that would force you to create combined categories like "Potato-Healthy" or "Tomato-Early_Blight" (resulting in 6 separate classes), multilabel classification allows the model to learn each dimension independently. This means better generalization, more flexible predictions, and the ability to easily extend your system - adding a new crop type or health condition doesn't exponentially increase your class combinations.
Creating and Populating Your Project
To get started on the Nexus homepage, click Create Project and select Multilabel Classification as your project type. This ensures the platform configures the appropriate annotation tools and training pipeline for multilabel workflows. Once your project is created, navigate to the dataset page where you can upload your leaf images in bulk. Nexus supports various image and video formats, and allows you to upload assets directly from your device or sync them from external cloud storage buckets like AWS S3 Bucket and Azure Blob Storage for seamless integration with your existing data infrastructure.
Annotating Your Images
After uploading your assets, it's time to begin labeling. When you first enter the annotator, you'll be prompted to create tags that will be used throughout your annotation workflow. For our agricultural leaf health example, we'll create five distinct tags:
- Potato_Leaf
- Tomato_Leaf
- Healthy
- Early_Blight
- Late_Blight
Notice how we've organized our tags to represent two independent dimensions: crop type and health status. This structure allows the model to learn these attributes separately, which is the core advantage of multilabel classification.
To annotate an image:
- Click on the image you wish to label
- Select the first applicable tag from your tag list
- Press A (or click "Assign") to apply the tag to the image
- Repeat steps 2-3 for each additional tag that applies to the image
With multiclass classification, you would just assign one class to an image like the image below, such as "Potato_Leaf_Late_Blight".
.webp)
For multilabel classification, you can assign more than one tag to an image. For example, if you're looking at an image showing a potato leaf affected by late blight, you would assign both Potato_Leaf AND Late_Blight tags to the same image. Unlike single-class annotation where you'd select only one category, multilabel classification empowers you to capture the complete context of each image through multiple simultaneous labels.
.webp)
As you annotate, pay attention to ensuring every image receives at least one tag from each dimension (crop type and health status) to maintain consistency in your dataset. This helps the model learn structured relationships between label categories.
If annotation at scale becomes time-consuming, Datature also offers intelligent tools to speed up your labelling process, and external labeling services with experienced annotators who can handle your multilabel classification projects efficiently and accurately.
Creating Your Training Workflow
Once you've successfully annotated your dataset, it's time to create your training workflow. Navigate to the Workflow page on Nexus, where you can configure your model architecture and training parameters. For this agricultural leaf health example, we'll be using HGNetV2, a lightweight yet powerful backbone architecture that's well-suited for multilabel classification tasks.
To set up your workflow:
- Add a Dataset Block - This connects your annotated images to the training pipeline
- Configure Augmentations - Apply transformations like rotation, brightness adjustment, and flipping to help your model generalize better across different lighting conditions and leaf orientations commonly found in field conditions
- Select the Model Block - Choose HGNetV2 B0 with an input image size of 320x320 pixels, which balances accuracy and inference speed for agricultural monitoring applications
For our training configuration, we'll use:
- Batch size: 10 images per training step
- Training duration: 5000 steps
These parameters provide sufficient iterations for the model to learn the independent relationships between crop types and health conditions without overfitting on our dataset.
Observing The Training
Once you click Start Training, you'll be redirected to the Training Run page where you can monitor live progress as your model learns. This is where the benefits of Nexus's real-time monitoring become apparent.
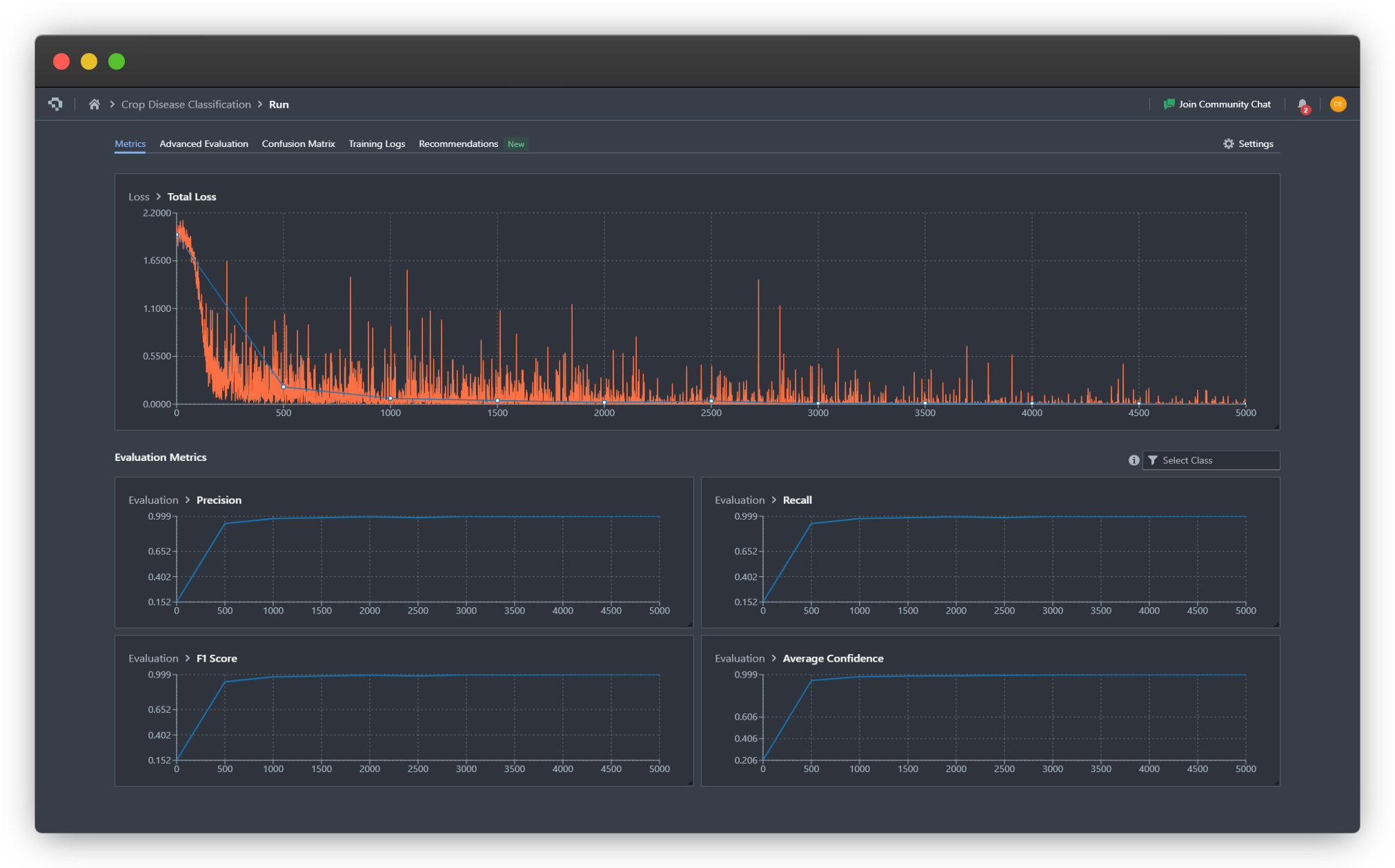
For our leaf health classifier, we can observe that the model converges around step 1000, as indicated by the stabilization of the loss curves and the F1 metric reaching a plateau. The binary cross entropy with logits loss (total loss) decreases steadily across both dimensions (crop type and health status), showing that the model is successfully learning to make independent predictions for each label category.
We also observe that the multilabel classification training seems to converge to a successful result more quickly, achieving higher precision, recall, and F1 score by step 500 and 1000, confirming our hypothesis that multilabel classification simplifies learning of labels and inter-label relationships.
Key metrics to watch:
- Training Loss - Should decrease steadily and stabilize
- Validation Loss - Should follow a similar pattern without diverging (which would indicate overfitting)
- Per-Label F1 Scores - Track how well the model predicts each individual label
Advanced Evaluation
For deeper insights into your model's performance, navigate to the Advanced Evaluation tab. Here you can examine predictions at specific checkpoints throughout training.
In our agricultural example, at step 500, the model was already capable of correctly classifying both the crop type and health status for most validation images, though with moderate confidence levels. By step 1000 and beyond, the model's confidence scores rose to nearly 100% for correctly labeled samples, indicating strong learned representations for both label dimensions.
This progressive improvement demonstrates how multilabel models learn multiple concepts simultaneously - the model didn't need to master crop identification before learning health classification, but rather developed understanding of both dimensions in parallel.

Co-occurance Matrix
The Co-occurrence Matrix tab provides crucial insights into whether your model has learned the true relationships between different tags - specifically, which labels are independent versus mutually exclusive.
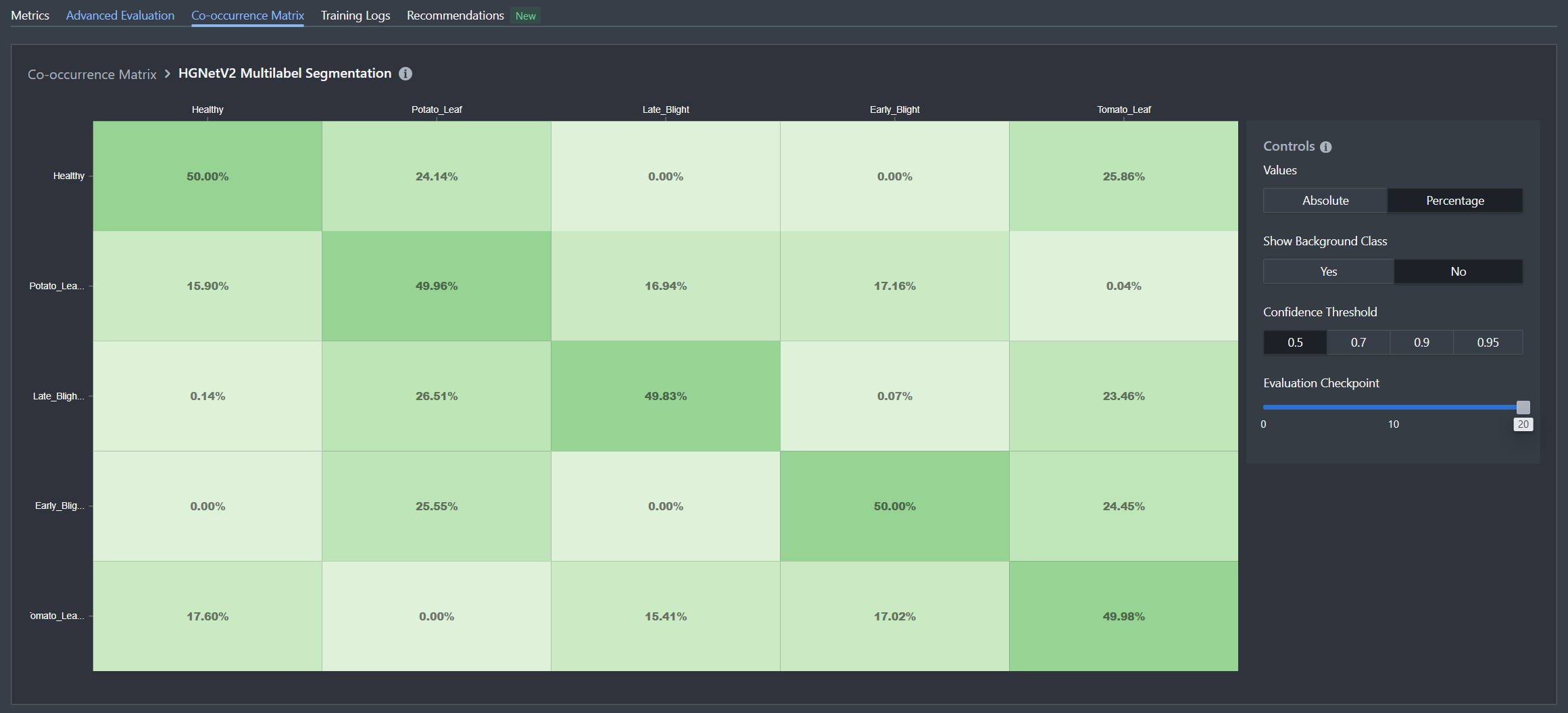
In our leaf health dataset, we can observe clear patterns that validate the model's understanding:
Mutual Exclusivity Within Dimensions:
- When the model predicts Potato_Leaf, it almost never concurrently predicts Tomato_Leaf (and vice versa)
- Similarly, health status labels (Healthy, Early_Blight, Late_Blight) rarely appear together on the same image
Independence Across Dimensions:
- Crop type predictions and health status predictions occur together freely, showing the model understands these are independent attributes
- For example, Potato_Leaf appears with all three health statuses in the predicted combinations
This matrix confirms that the model has correctly learned the underlying structure of your labeling scheme: crop types are mutually exclusive, health conditions are mutually exclusive, but crop types and health conditions are independent of each other. This is exactly the behavior we want from a multilabel classifier tackling multi-dimensional classification problems.
If you notice unexpected co-occurrence patterns - such as certain label combinations that should appear together but don't - this could indicate either dataset imbalance or the need for additional training examples featuring those specific combinations.
What Happens Next?
Once you're satisfied with your trained multilabel classification model's performance, you have several options for putting it into production. Nexus provides flexible deployment pathways designed to fit different agricultural monitoring workflows and infrastructure requirements.
Export Your Model
You can export your trained model in any of our supported formats, allowing you to integrate it into existing systems or custom applications. Nexus also supports pruning and quantization during export to optimize your model for resource-constrained environments.
Deploy with Datature
Alternatively, deploy your model directly through Datature's deployment solutions without writing additional code:
Datature Cloud - Host your model in the cloud and access it via API for real-time leaf health analysis from any location. Ideal for centralized monitoring systems or integration with farm management software.
Datature Outpost - Deploy on-premises for scenarios requiring data privacy, offline operation, or low-latency inference. Perfect for greenhouse monitoring systems or agricultural research facilities.
Local Devices - Run your model directly on edge devices like cameras, drones, or handheld scanners for immediate in-field diagnostics without internet connectivity.
For our agricultural leaf health example, you might deploy the model to a mobile device that farmers can use directly in the field, instantly identifying both the crop type and any disease present to enable rapid intervention.
Ready to get started? Explore our deployment options and bring your multilabel classification model to life.
Putting It All Together - Why Multilabel Classification Matters
Multilabel classification represents a significant expansion of Datature's capabilities, enabling you to build more sophisticated computer vision systems that mirror the complexity of real-world scenarios. By allowing models to recognize multiple attributes, properties, or objects within a single image, you can create solutions that provide richer, more actionable insights from your visual data.
Whether you're building content moderation systems that need to flag multiple policy violations, e-commerce platforms that tag products with multiple attributes, or medical imaging tools that identify multiple conditions, multilabel classification provides the flexibility your application demands.
This feature is now live on the platform. We encourage you to explore how multilabel classification can enhance your computer vision workflows and unlock new possibilities for your projects.
Explore Further with Our Developer Roadmap
Have questions or feedback about the Multilabel Classification feature? Join our Community Slack to connect with other developers, share insights, or get support from our team.



.png)
.png)